Overview
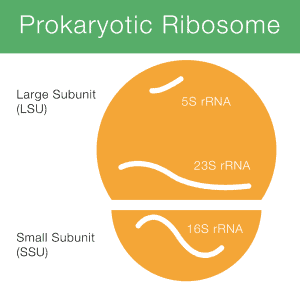
16S rRNA stands for 16S ribosomal ribonucleic acid (rRNA), where S (Svedberg) is a unit of measurement (sedimentation rate). This rRNA is an important constituent of the small subunit (SSU) of prokaryotic ribosomes as well as mitochondria and chloroplasts. Figure 1 displays how the 16S rRNA (shortly 16S) is involved in a prokaryotic ribosome.
DNAsegment coding for rRNA is called either rRNA gene or rDNA. For the purposes of sequencing, sequence information is obtained from the 16S gene because DNA is much easier to process and sequence than RNA.
Why 16S rRNA was chosen as DNA barcode molecule for taxonomy and ecology
- To be used as a DNA barcode, a gene should have the following characteristics.
It should be ubiquitously present. Otherwise, we cannot include all organisms. All members of Bacteria and Archaea are known to have 16S gene. - It should contain sufficient phylogenetic information. 16S is about 1,500 bp long, which is not too short or long.
- The genetic variation within 16S gene found among prokaryotes is adequate to be used in the phylogenetic analysis for the broad taxonomic ranges. It is successfully used to infer phylogenetic relationship among phyla while also used in the comparison among species in the same genus.
- It should be easily amplified by PCR. 16S gene has multiple conserved regions that can be used as the priming sites. This becomes a significant advantage for NGS-based short read sequencing.
- After many years of international collaboration, we have 16S sequence database holding almost all known species of Bacteria and Archaea. By searching 16S sequence against these databases, anyone, even without knowledge of serious taxonomy, can identify newly isolated strains.
Variable regions of 16S and PCR primers to amplify them
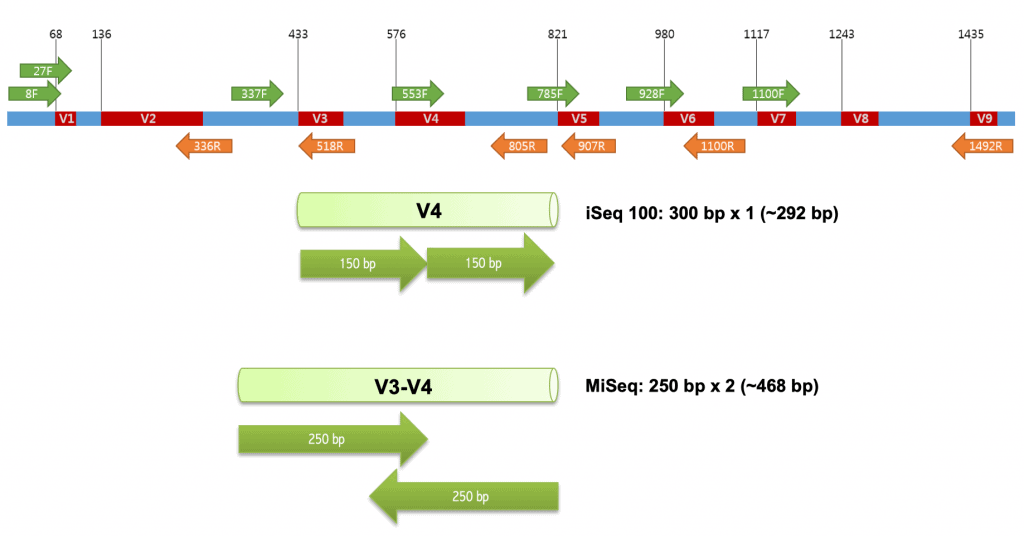
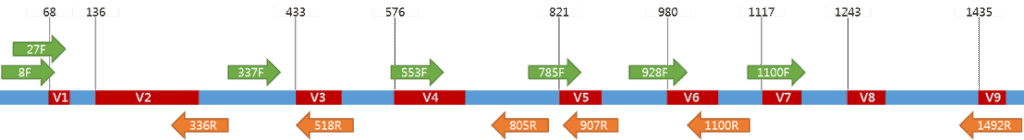
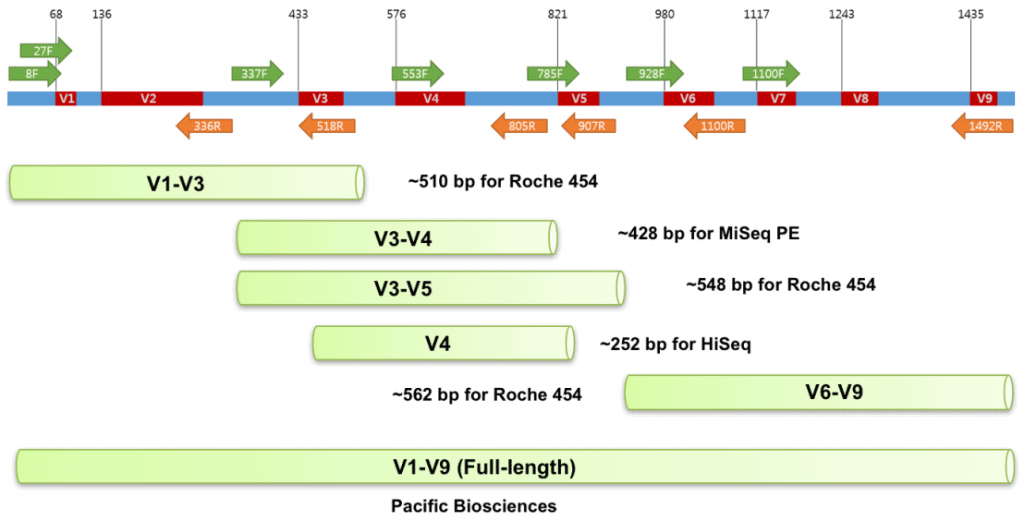
Sequence variation among bacterial 16S is known to be not uniformly distributed. Nine hypervariable regions were identified among Bacteria, which are named V1 to V9.
| Name of primer F=forward,R=reverse |
Sequence |
|---|---|
| 8F | AGAGTTTGATCCTGGCTCAG |
| 27F | AGAGTTTGATCMTGGCTCAG |
| 336R | ACTGCTGCSYCCCGTAGGAGTCT |
| 337F | GACTCCTACGGGAGGCWGCAG |
| 337F | GACTCCTACGGGAGGCWGCAG |
| 341F | CCTACGGGNGGCWGCAG |
| 515FB | GTGYCAGCMGCCGCGGTAA |
| 518R | GTATTACCGCGGCTGCTGG |
| 533F | GTGCCAGCMGCCGCGGTAA |
| 785F | GGATTAGATACCCTGGTA |
| 805R | GACTACHVGGGTATCTAATCC |
| 806RB | GGACTACNVGGGTWTCTAAT |
| 907R | CCGTCAATTCCTTTRAGTTT |
| 928F | TAAAACTYAAAKGAATTGACGGG |
| 1100F | YAACGAGCGCAACCC |
| 1100R | GGGTTGCGCTCGTTG |
| 1492R | CGGTTACCTTGTTACGACTT |
The full-length 16S gene is usually amplified by the pair of primers 27F and 1492R, followed by Sanger DNA sequencing. To obtain accurate sequence, both DNA strands should be sequenced using multiple primers given in the above table.
Application of NGS to bacterial community analysis
NGS is right for elucidating bacterial community structure, as it eliminates the requirement of tedious E. coli cloning and allows high throughput DNA sequencing. Because different lengths of DNA are sequenced by various NGS platforms, a suitable pair of PCR primers should be used. Three types of NGS sequencing are possible:
- Single-end sequencing: DNA sequence information is obtained from only one end of the PCR amplicon. Roche 454, Ion Torrents
- Paired-end sequencing: DNA sequence information is obtained from both ends of the PCR amplicon. Two sequences should be overlapped by a sufficnet length to combine to become a single sequence. Illumina
- Circular consensus sequencing (cca): Pacific Biosciences’ SMRT technology provides very long single molecule reads ranging >10K bp. Because of the circular-shaped library used, the same 16S amplicon can be sequenced multiple times to produce accurate consensus sequence.
Recommended sequencing formats
| NGS systems | 16S region | PCR primers | Estimated insert size (E. coli) | Sequencing format |
|---|---|---|---|---|
| Illumina MiSeq | V3V4 | 341F & 805R | 428 bp | 250 x 2 |
| Illumina iSeq 100 [Learn more] | V4 | 515FB & 806RB | 252 bp | 300 x 1 |
| Illumina HiSeq | V4 | 515FB & 806RB | 252 bp | 150 x 2 |
Last updated Feb. 20, 2019