What is a species?
A species is often defined as a group of individuals that can actually or potentially interbreed in nature. In practice, this concept cannot be easily applied to any species. For example, you may not know with confidence if two insects that you encountered in a rain forest belong to the same species, as it can only be confirmed by breeding them under natural conditions (and you observe it!). Getting into the species concept of animals is this difficult. What would that be like for our invisible, but intimate strangers, Bacteria?
It would be best if I gave you an example to explain how we actually recognize bacterial species.
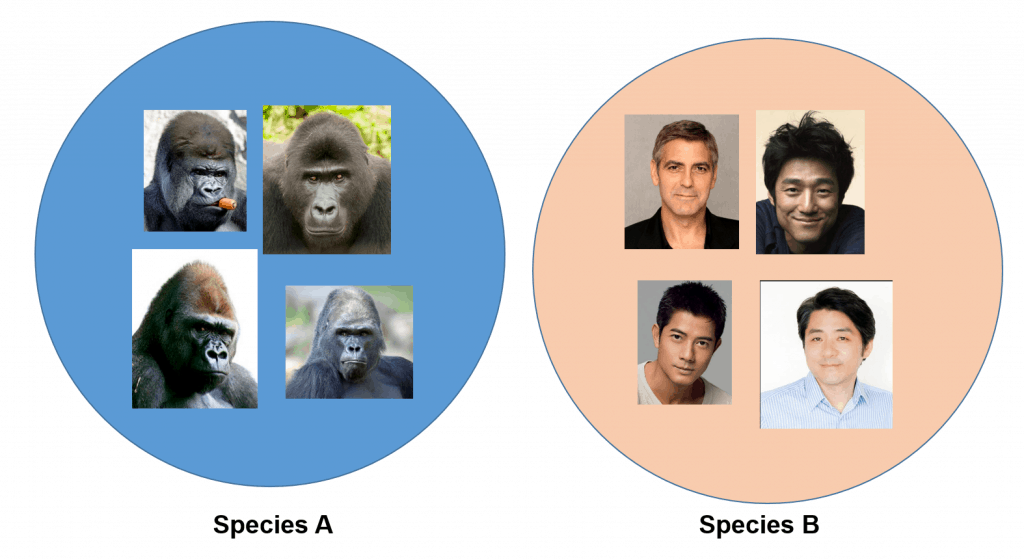
The pictures below show individuals belonging to two different species:
You do not need to be a taxonomist to classify them into two different species (..at least I hope not!)
We will not try to confirm this classification by interbreeding them, but our visual observation provides sufficient evidence for such a task (e.g. presence of extensive facial hair and jaw shape, etc.).
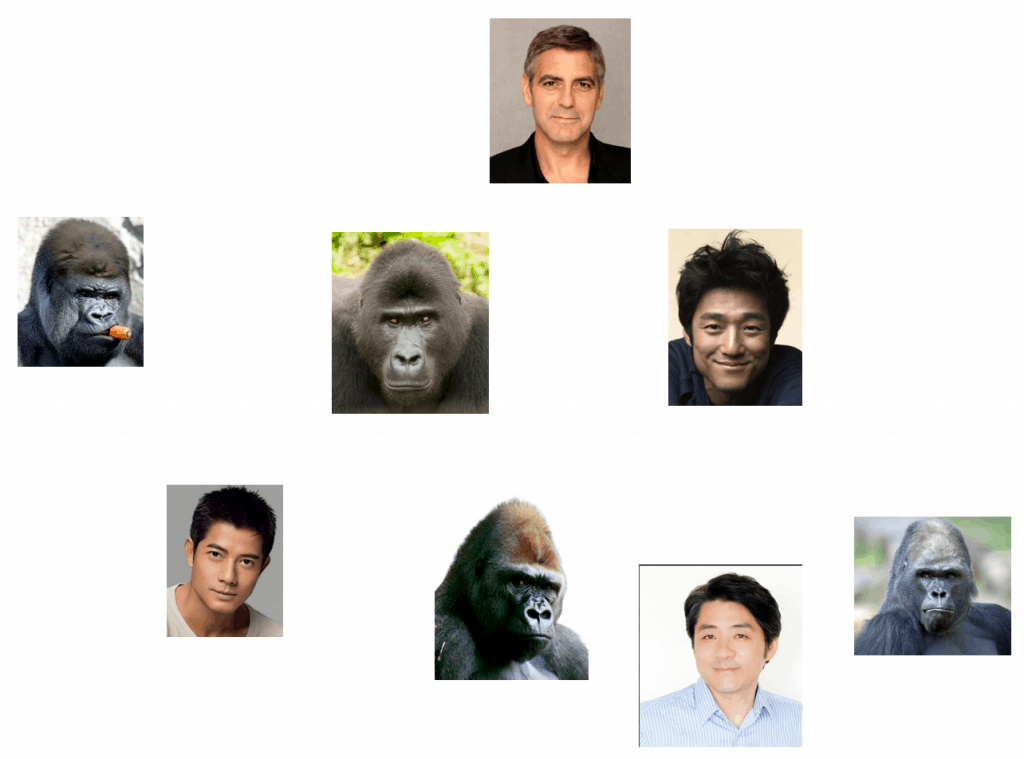
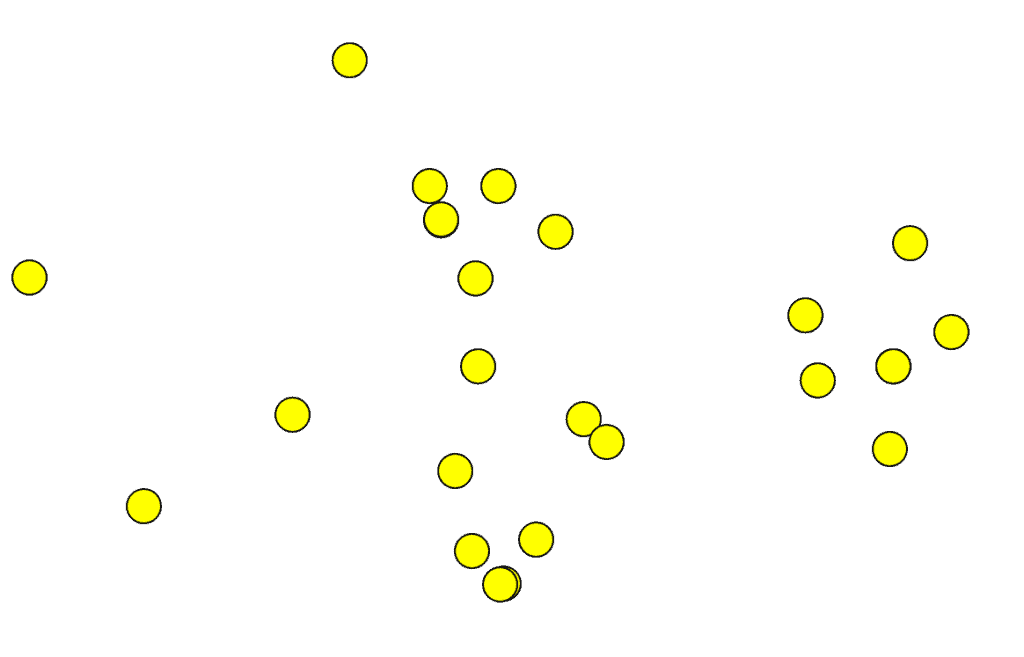
Now, let’s try to classify bacteria depicted in the following pictures using your expertise:
Can you tell me how many species are there in the above figure? Probably not. Even a king of bacterial taxonomy, if there is such a thing, could not classify these correctly just by observing the external shape of cells. This is why bacterial classification is a difficult task. And, the way we classify depends on how we define the species in bacterial world. So, the question remains, “how can we define bacterial species?”
Bacterial Species Concept
Many people think that science is ruled by some sort of governing body. This is partly true for bacterial taxonomy. There is a body called “International Committee on Systematics of Prokaryotes (ICSP)” which plays a role similar to the United Nations in international politics. Even though ICSP provides recommendation reports from time to time, it cannot formally setup the definition of bacterial species. It is rather decided by community efforts or consensus. At present, the most widely accepted species concept is called “Phylo-phenetic species concept” (Rosselló-Mora & Amann, 2001).
“A monophyletic and genomically coherent cluster of individual organisms that show a high degree of overall similarity in many independent characteristics, and is diagnosable by a discriminative phenotypic property.”
There are several terms that I need to explain further:
- “A monophyletic cluster” contains a group of bacterial strains that shared a most recent common ancestor.
- Bacterial strains forming a “genomically coherent cluster” show similar genome sequences, thereby likely similar phenotypes. Just like you and your brother look more similar than you do to your friends as you and your brother share more genome sequence similarities.
- “Overall similarity in many independent characteristics” means that two strains share phenotypic properties that are not related or interconnected. For example, the height and weight of humans correlate therefore these characteristics are not independent.
- “Diagnosable by a discriminative phenotypic property” means that two species should be differentiated by phenotypic characteristics (such as biochemical tests, morphology and physiology). If two species can only be differentiated by means of genome or 16S rRNA sequences, this two should not be recognized as different species. This condition ensures that people who do not have molecular techniques or facility can still carry out identification. However, I think, as more people, if not most, are moving towards molecular diagnostic methods (PCR, DNA sequencing, Microarray even NGS) over traditional biochemical tests, this condition can be omitted in near future (I hope). Furthermore, microbiologists, other than taxonomists, are already using solely molecular concept of bacterial species as molecular OTU.
The Practical Bacterial Species Concept
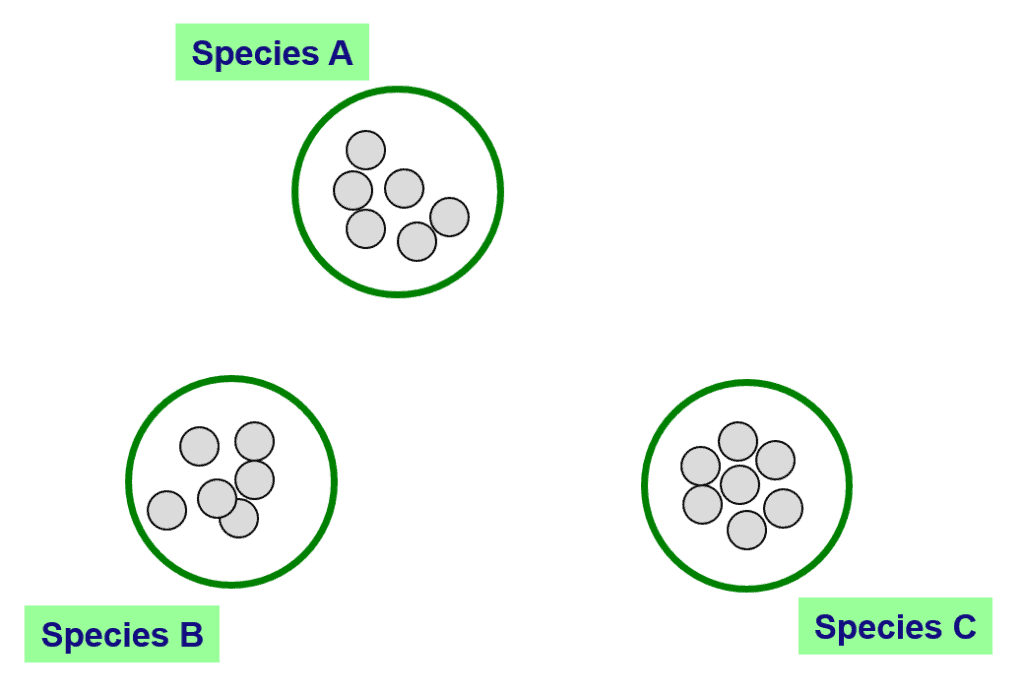
“Phylo-phenetic species concept” sounds very solid. However, its application can be very tricky. Again, let me explain by some examples.
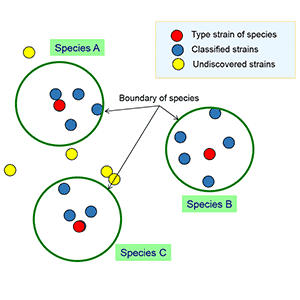
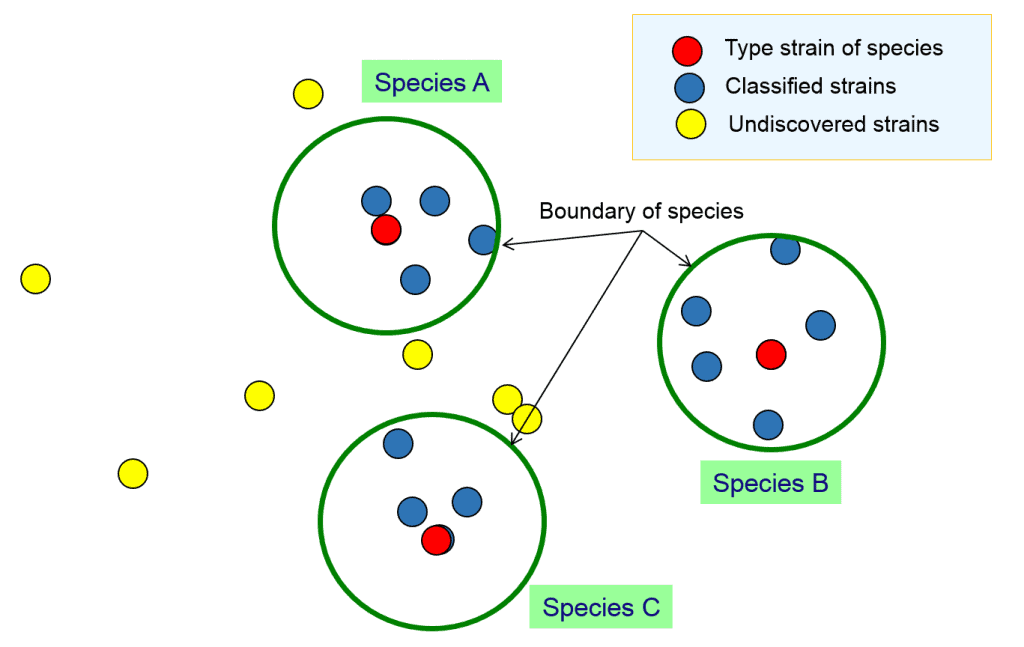
In the above figure, you can see 3 clearly differentiated clusters that can be confidently called species A, B and C. How about the below case?
Well to me, the clear divisions of clusters are not apparent. However, we still need to classify and name the species (as we want to call them by names not bacteria X or 110982). To achieve this, bacterial taxonomists have introduced a concept of “type strain”. A type strain is a live strain that can serve the center of a species and regarded as the representative of a species. When multiple strains are discovered for a single species, we can choose a likely representative strain as the type strain. In practice, most of bacteria species are described with only one or two strains, the type strain of a species is often the strain which was first discovered. What I am trying to say is that “type strain” may not be very “typical” for a given species! For example, the type strain of Escherichia coli does not kill you, but other strains, such as O157 strains, can kill you easily.
Once the type strain is decided and deposited to the institutions called culture collections, it cannot be easily changed to other strain since the stability is very important virtue of taxonomy!
Let’s assume that a team of taxonomists carried out research to classify the strains in the previous figure and come up with the following result:
Here, the team found 3 species and designated 3 type strains for each. As a species is a coherent group of bacterial strains, we should employ the same measure of “coherence” or “similarity” for all 3 species. We need to define the followings to be “objective” for this classification process.
- How the similarity can be defined? (A method of measuring similarity/distance)
- How similar should be between the type strain of species A and a strain belonging to species A. (The cutoff of species boundary)
Defining the above two criteria has been major challenge for modern bacterial taxonomy. In 1987, major players in the field of bacterial taxonomy have gathered in Paris to try to come up with an objective and stable criteria for future classification and identification of Bacteria. They foresaw that genome data (genotypic) are superior to phenotypic data (physiology and biochemisty), but sequencing of genome was not readily available until 1995. However, at that time, there were other molecular method, called DNA-DNA hybridization (DDH), to measure the degree of hybridization of genomes in solutions. If two genomes hybridize well, they should share similar nucleotide sequences.
DDH provides overall, albeit indirect, measure of genomic similarity between two strains, and serves well as a surrogate for genome sequence comparison. In a seminal paper, Wayne and other taxonomists recommend DDH as the method for defining bacterial species and 70% relatedness as cutoff for the species boundary (Wayne et al., 1987). This Wyane et al. paper has been cited over 4,000 times which means that this proposal was well received. In conclusion, if a strain belongs to a species, it should show 70% or higher DDH relatedness value to the type strain of that species.
Genome sequence-based species concept
Thanks to the introduction of next generation sequencing (NGS), bacterial sequencing is now cheap enough and readily available to many researchers. I believe that genome sequence information is the best you can get for any taxonomic work that can eliminate the needs for many tedious and unreliable experimental taxonomic methods. Of course, it can replace the notoriously erroneous DDH in the definition of bacterial species. “Overall Genome Related Index (OGRI)” is a term for any computational method to calculate similarity between two genome sequences, first coined by Fred Rainey and myself in 2014. There are many different algorithms that can be used for comparing two strains, Average Nucleotide Identity (ANI) has been most widely accepted. The generally accepted cutoff value for the species boundary is about 95~96% ANI. Here I recommend you the OrthoANI algorithm, an improved version of ANI, instead of the original ANI. (More about OrthoANI).
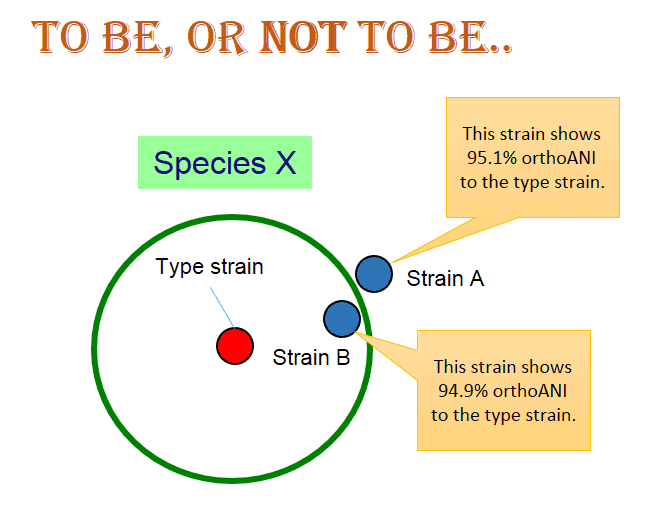
For both ANI and OrthoANI, about 95~96% is the cutoff. Does this mean this cutoff is really a clear and sharp one that can be used without exception? Let’s consider the following case:
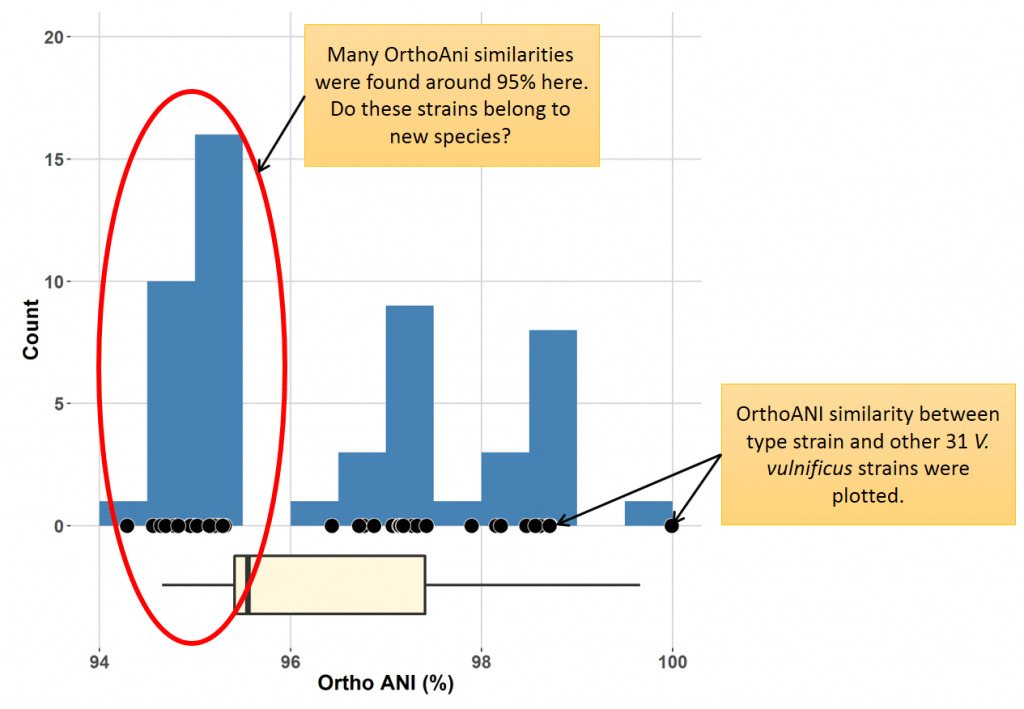
Two strains show 95.1 and 94.9% OrthoANI, respectively, to the type strain of “species X”. Does this mean that strain A belongs to “species X” and stain B does not? You may think that I made this case up and it is not a probable case? The below is the real case of Vibrio vulnificus, a notorious pathogen from sea water.
Here is a chart in which 31 V. vulnificus strains were examined for OrthoANI against to the type strain of the species. Many strains show OrthoANI values around 95%.
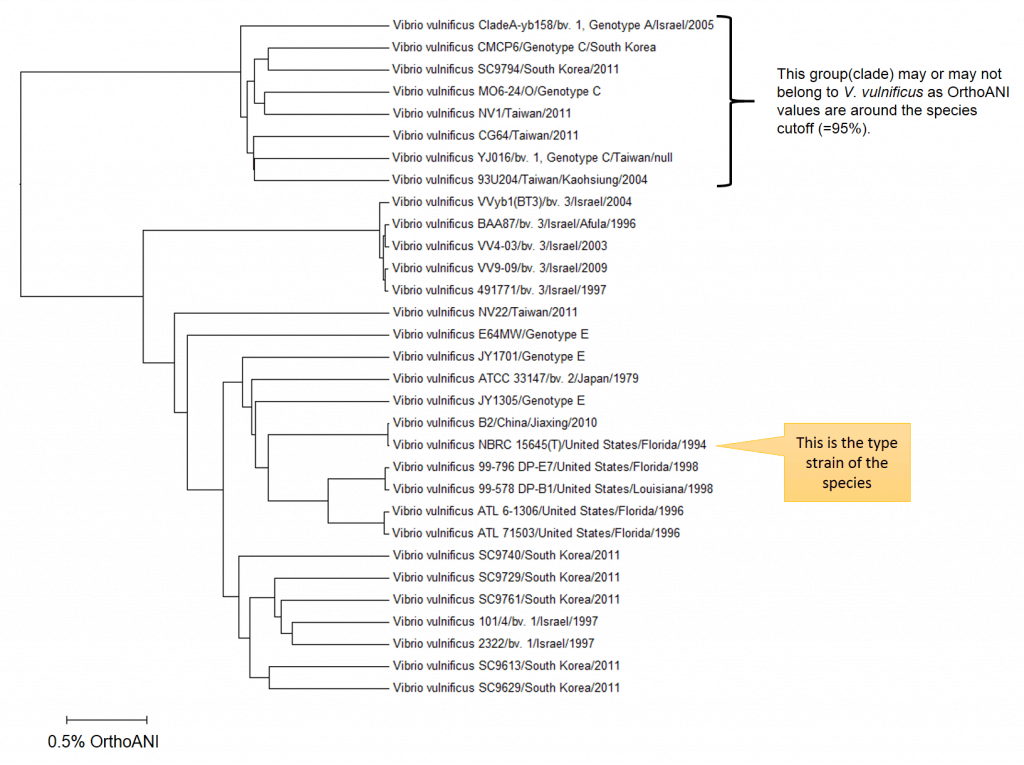
When we look at the above dendrogram explaining overall taxonomic structure within the V. vulnificus , these strains may belong to the different species. However, OrthoANI values between the authentic V. vulnificus group (containing type strain) and the outlier group are around the proposed cutoff, i.e. 95%, therefore the decision is not a straight-forward one. In my opinion, two groups can be either different species or at least different subspecies. Anyhow, it is up to taxonomists who will work on the further evidence and draw the final conclusion. Meanwhile, I can only tell you that V. vulnificus is not a really one genomically coherent group.
Take home messages:
- Bacterial species is defined as “genomically coherent group of organisms”.
- A species must have a type strain that is live. Anyone can obtain this strain for taxonomic study.
- If a strain belong to a species, ORGI should be within the cutoff of species boundary. If you use OrthoANI , it will be about 95%.
- If your isolate is a new species by the condition (3) and you want to describe it as new species, you need to carry out additional taxonomic research (mostly phenotypic characterizations such as biochemical tests etc.). Even though we all know that you have new species by genome sequence comparison, this allows us to understand more about the biology of species. Genome sequence tells us a lot, but not sufficient enough for us to understand their physiology and life style.
References
- Rossello-Mora, R. & Amann, R. The species concept for prokaryotes. FEMS Microbiol Rev 25, 39-67 (2001).
- Wayne, L.G. et al. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37, 463-464 (1987).
By Jon Jongsik Chun (CEO of CJ Bioscience, Inc. & Professor at Seoul National Univ.)