All potential orthologous protein-coding genes (=CDSs) are clustered into non-redundant gene sets after pan-genome calculation to generate “Pan-genome Orthologous Groups (POGs)”.
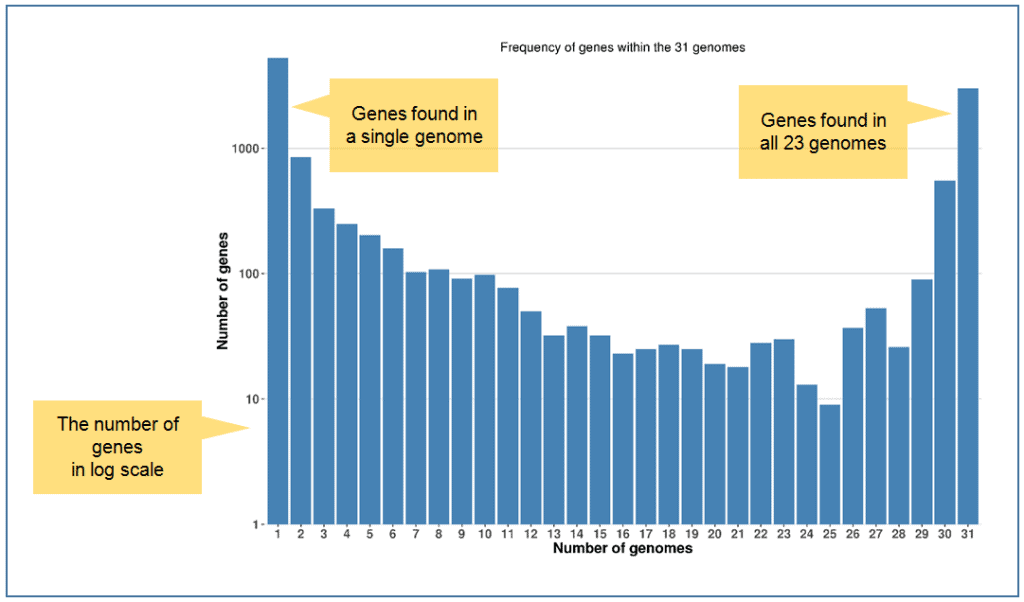
Obviously, a core part of the genome containing essential or house-keeping genes are found within all genomes, and less important genes are found less frequently. Some genes are detected only in a single genome. A “gene frequency plot” gives a general overview of the frequency of genes within a whole genome set. A typical plot will show a U-shaped plot where most genes are detected either as all genomes or a single genome.
In the following example, 31 Vibrio vulnificus genomes were analyzed from which 13,220 non-redundant genes were found from a total of 144,931 genes. Please note that the below figure shows the number of genes in log-scale.
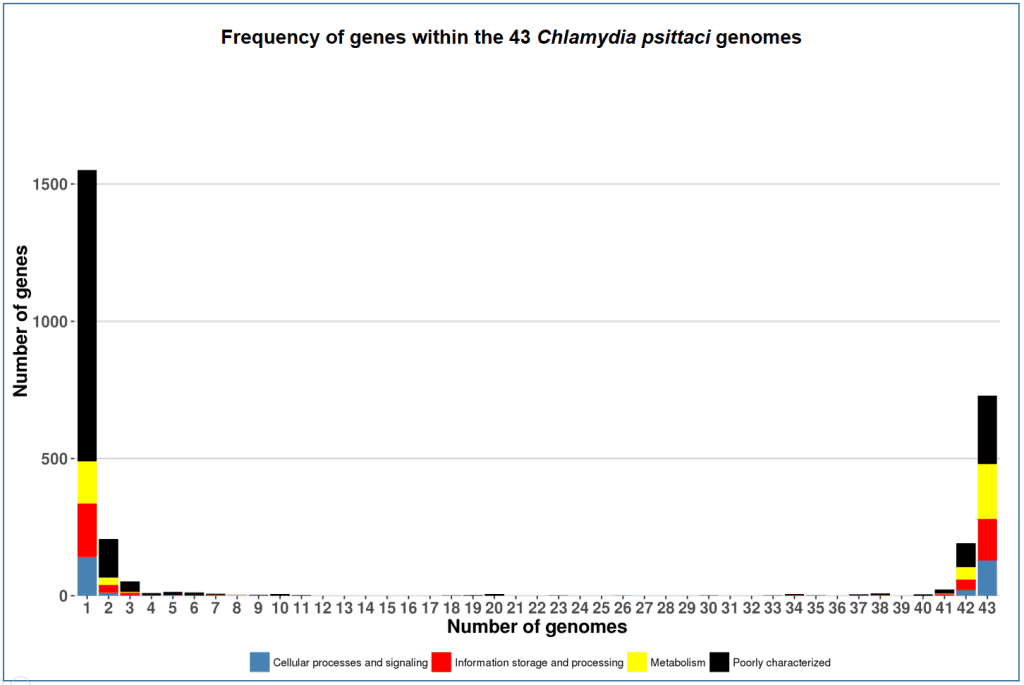
The below figure visualizes the same data as the above except that the genes are classified into 4 major functional categories, and the number of genes that are displayed are NOT in log-scale.
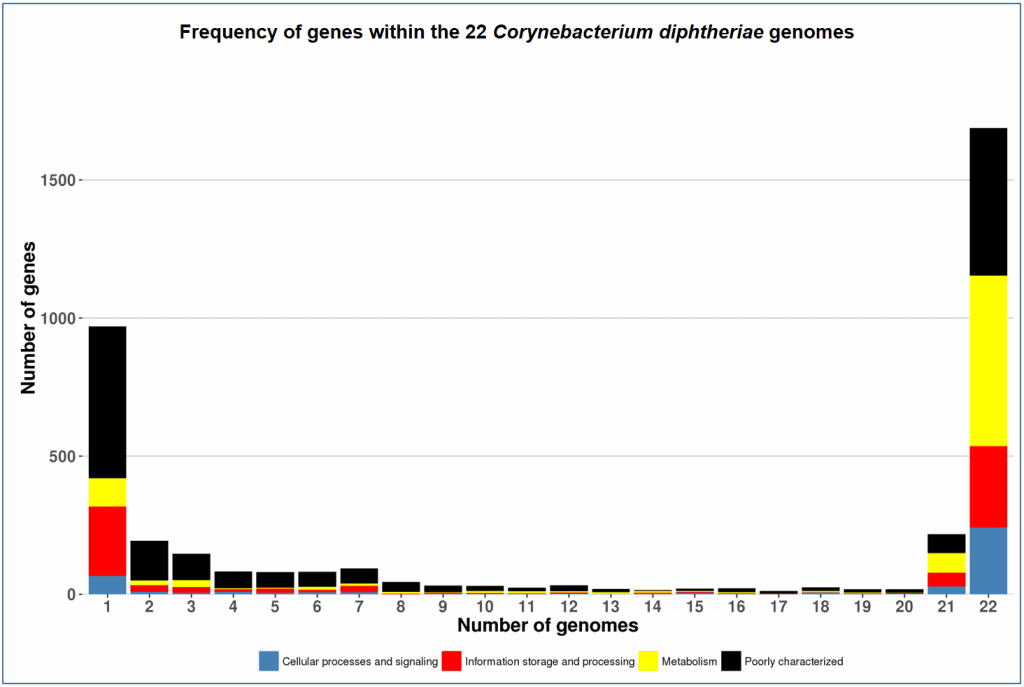
Here are more pan-genome examples of other species. Can you tell how and why the shapes of the charts are different?
- Chlamydia psittaci (obligate parasitic bacterium with small genome)
- Corynebacterium diphtheriae (Pathogenic bacterium belonging to the phylum Actinobacteria)
This type of figures has been used in many publications including:
- Lefebure, T. & Stanhope, M.J. Evolution of the core and pan-genome of Streptococcus: positive selection, recombination, and genome composition. Genome Biol 8, R71 (2007).
- Touchon, M. et al. Organised genome dynamics in the Escherichia coli species results in highly diverse adaptive paths. PLoS Genet 5, e1000344 (2009).
Updated on April 28th, 2016 (EK)