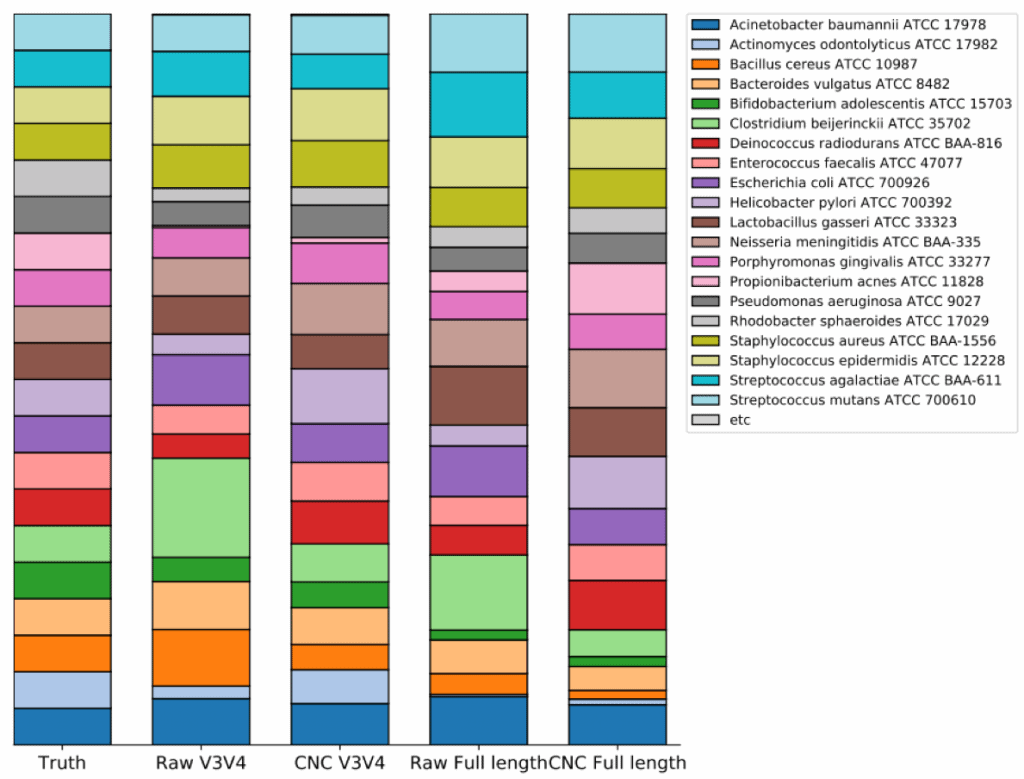
Several standards products are available to validate 16S microbiome sequencing procedures. They are usually a mixture of genomic DNAs or whole cells from different species whose composition is precisely known. At present, EzBioCloud can be used along with these products including one sold by the American Type Culture Collection (ATCC). In this tutorial, we offer a step-by-step guide to help you use this product.
What to prepare in advance
You need NGS data to upload:
You need NGS data file(s) from 16S amplicon sequencing from ATCC Microbiome Standards product. If it is from Illumina (paired-end or single-end), it should be in FASTQ format. For Pacific Biosciences, it should be in FASTA format (after ccs processing). If you don’t have data, don’t worry. We provide an example file you can download from here . These ATCC Microbiome Standards were sequenced by CJ Bioscience.
You need an account at EzBioCloud:
You will also need an account at www.EzBioCloud.com. If you don’t have one already, please register at //www.ezbiocloud.net/signup. It is free but requires you use an institutional email account (not gmail etc.). Once you are logged into EzBioCloud.com, you will also need to sign up for a free trial of the Microbiome Taxonomic Profiling (MTP) service. Please visit here to request your free trial. Your free trial account will let you analyze and compare up to 20 microbiome samples, each containing up to 100,000 reads.
Once you have a EzBioCloud account with a free trial for the MTP service activated, and data to upload, you are ready to go.
What to do in this tutorial
We will do the following in this tutorial. If you want to know further about any specific items, please search within help.ezbiocloud.net.
Please follow the below tutorial. If you have any question, please contact us at bs.ngs@cj.net.
The EzBioCloud team / Last edited on May 22, 2018