Human Microbiome Project funded by US National Institutes of Health (NIH) resulted in the taxonomic profiling data of microbiomes on 19 body sites of 242 healthy human subjects. Here we processed 8,048 16S microbiome taxonomic profiles (MTPs; Learn more about MTP) using EzBioCloud 16S database and EzBioCloud MTP pipeline. This document will guide you with how to access individual data. The average taxonomic coverage at the species level is >96% [Learn more].
- Login to www.ezbiocloud.net (It’s free to create an account).
- Open [Apps] at the top-right corner of the screen.
- Choose [MTP(Microbiome Taxonomic Profiling)] to go to the MTP main page.
- Open [+ Public Database] button to open “EzBioCloud Microbiome Database” which contains tons of 16S microbiome data that can be imported into your account.
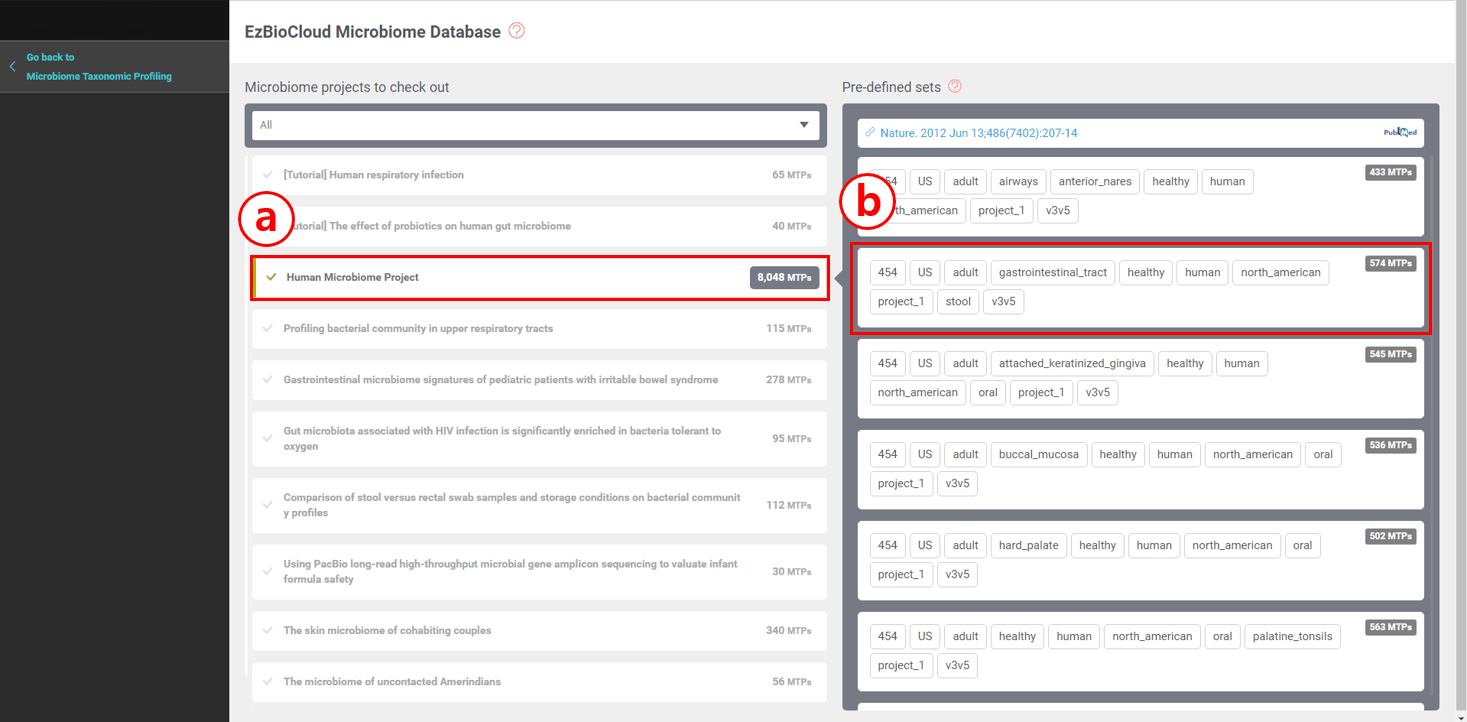
Now, you are in the EzBioCloud Microbiome Database.
- Human Microbiome Project (HMP) data are available here. Note that there are 8,048 MTPs (samples) which are divided into subsets with appropriate metadata tags. Remember that HMP data has a tag named “project_1”. You can select HMP data in the MTP workspace anytime using this tag.
- In this tutorial, I will choose “stool” data. By clicking here, 574 MTPs will be imported into my MTP workspace.
Select [Go back to my data] to come back to MTP workspace.
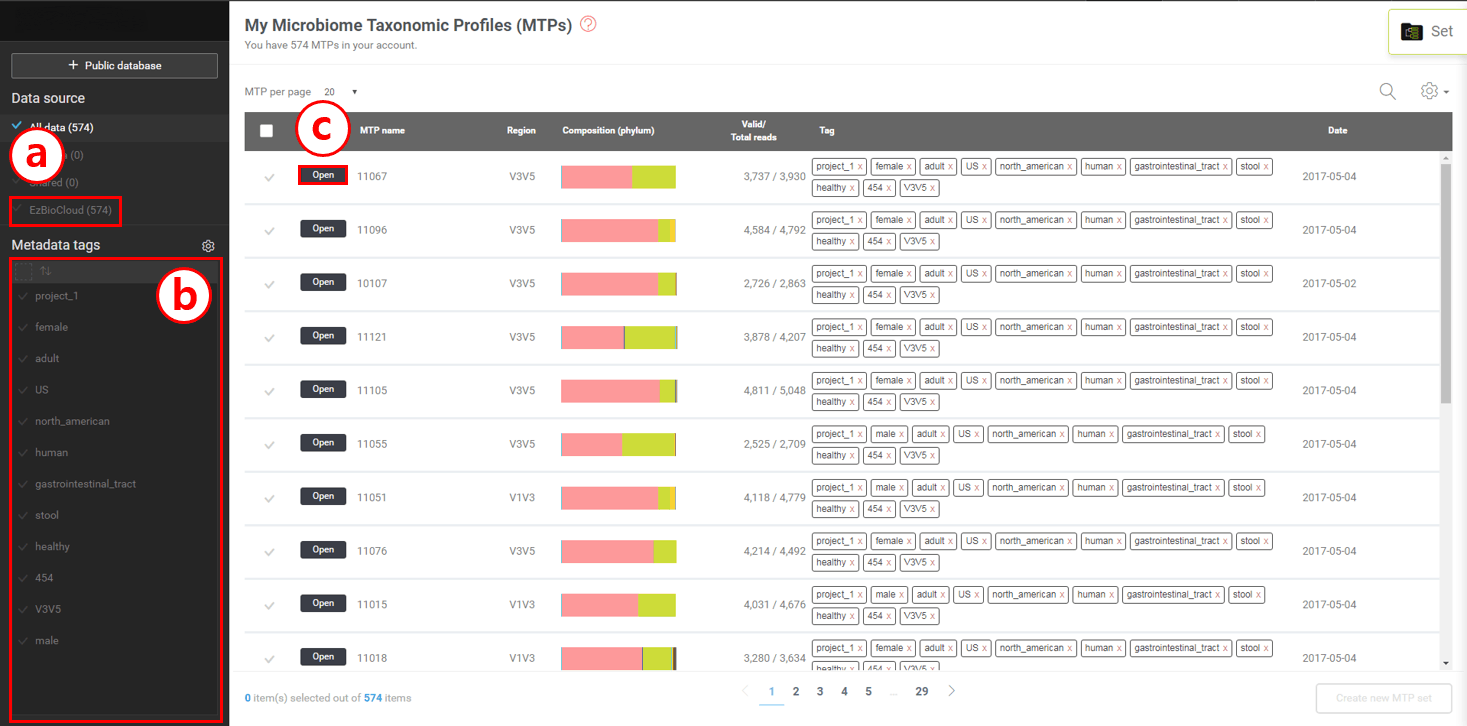
- Now you have 574 MTPs in your workspace.
- Metadata tags were imported as well for you to choose.
- Click [Open] to explore individual MTP from the HMP dataset.
Last updated by JC on Aug 27, 2017